201901 Clinical Genomics
Clinical Genomics Track
Zulip Chat
The Zulip Chat will be used to coordinate among participants during the connectathon:
Submitting WG/Project/Implementer Group
Justification
Genomic data are of increasing importance to clinical care and secondary analysis. After initial Feedback from the first ballot of the Clinical Genomics Implementation Guide, considerable updates have been made, and the recently balloted IG and profiles should be tested in the Connectathon for more feedback and trial use.
Proposed Track Lead
Gil Alterovitz, Patrick Werner
Expected participants
Gil Alterovitz, James Jones, Kevin Power ...
Roles
FHIR Client
Support the sending of the Sequence resource/genetics profiles operations: create, history, read, search and update.
FHIR Server
Support the receiving and processing of the Sequence resource/genetics profiles operations: create, history, read, search and update.
PGx CDS Service
PGx CDS Service will interact with EHR (via CDS Hooks), OMIC Data Store (for genomoic observations), and Terminology Server (for medication value sets).
OMIC Data Store
Will store genomic observations
EHR
Order entry of a medication triggers a “medication-prescribe” CDS hook which invokes the PGx CDS Service and provides it with a FHIR MedicationOrder instance.
Terminology Service
The PGx CDS Service will ping the terminology server to see if the ordered drug is in an “Actionable PGx Drug” value set.
Scenarios
Register a New Sequence and Observation
Register a New Genomics Report
Scenario 1 PGx (placeholder)
Scenario 2 Clinical Sequencing - Germline Testing
Scenario 3 Family Member History
Scenario 4 Clinical and Research Data Warehouses
Scenario 5 HLA Typing
Scenario 6 Comprehensive Pathology Report
PGx CDS Scenario
Please see detailed storyboard here: https://docs.google.com/document/d/1VqPrf6aaOB8RhSbAq2JYDxZYbDm-RT9DxA0u6-RxYo0/edit
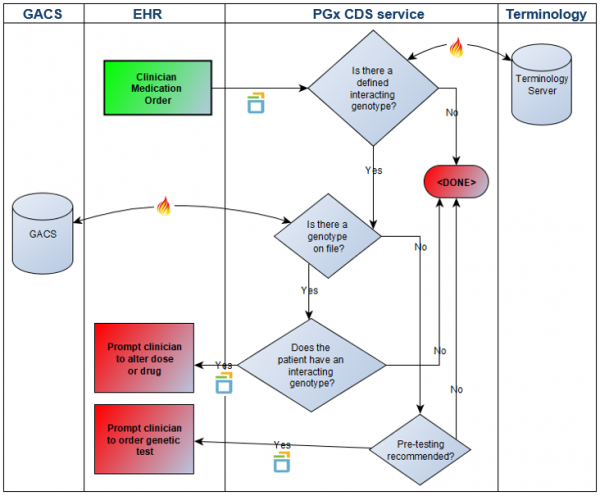
A swim-lane overview of the scenario is here:
- Action: [1] (EHR) Manually enter an order for "Imuran 50mg 1 tablet twice a day by mouth" into order entry system. This triggers a “medication-prescribe” CDS hook with a hook context of MedicationOrder which includes RxNorm code 197388 for azathioprine 50mg oral tablet; [2] (PGx CDS Service) Having been triggered by the “medication-prescribe” CDS hook, the PGx CDS Service now executes a decision support rule that [1] determines if the ordered drug has a known gene interaction; [2] determines, where there is a known drug-gene interaction, whether or not the patient has genetic test results on file; [3] determines, where there are genetic test results on file, if the patient has an interacting genotype; [4] determines, where there are not genetic test results on file, if the patient needs pre-testing. PGx CDS Service returns a CDS hooks “information card” back to the EHR, with appropriate recommendations.
- Precondition: Genomic observation(s) are available for some patients.
- Success Criteria: Retrieve relevant observations (on TPMT gene, on allelic state) where they exist.
- Bonus point: Search OMIC data store for different potential representations.
- Reference: Azathioprine Pharmacogenomics - CPIC recommendations [1]